Uncovering hidden protein structures

Using infrared (IR) spectrosocopy, researchers at the University of Konstanz were able to uncover the interaction between the p53 protein, a tumour suppressor that controls the cell cycle, and poly(ADP-ribose) (PAR) and deoxyribonucleic acid (DNA) at the molecular level. The nucleic acid-like biopolymer PAR serves as a cellular signal transmitter and helps to regulate protein activity. By studying the interaction between p53 and PAR, the researchers were able to learn more about molecular reactions to cellular stress in response to, for example, DNA damage, which represents a potential tumour risk.
Their basic research on the processes behind DNA damage is, on the one hand, paramount to understanding how cancer develops and how cells age. On the other hand, the innovative scientific approach is advancing the research carried out in their field. Their research results were published in issue 9 (21 May 2019) of the scientific journal Nucleic Acids Research by the Oxford University Press.
Innovative connection
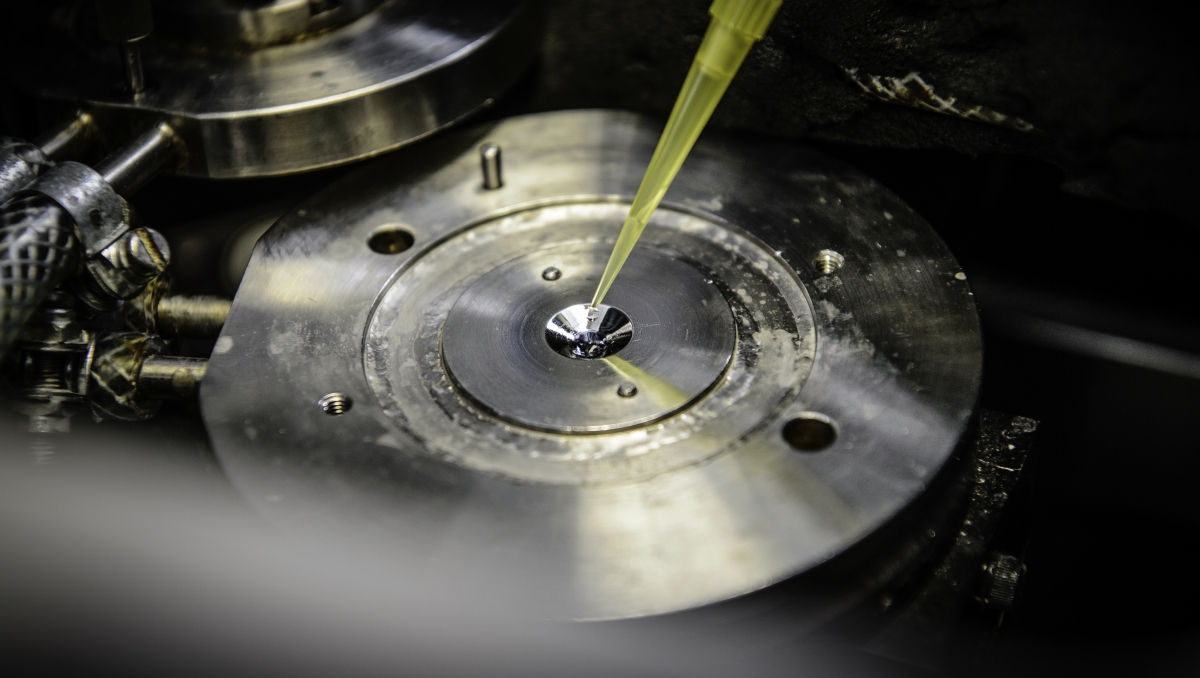
IR spectroscopy is a measurement technique used to investigate molecules. By exciting a molecule with infrared radiation, researchers can analyse the vibrational energy within the molecule, and thus closely track molecular processes. “IR spectroscopy is an established and widely used method in the chemical analysis of small molecules.
We applied this method in the present study in order to investigate the interaction between biological macromolecules that play a decisive role in the cellular stress response to DNA damage,” explains Karin Hauser, Professor of Biophysical Chemistry at the University of Konstanz.
“It was in fact interdisciplinary and research-oriented teaching at the University of Konstanz that provided the impulse for this research project at the interface between chemistry and biology. Staff internships played a pivotal role here. By joining more experienced researchers in the laboratories, master’s students in Biology, Chemistry, and the Life Science study programme were able to experience research first-hand."
Aswin Mangerich, lecturer in Toxicology and Biochemistry and member of the Molecular Toxicology research team headed by Professor Alexander Bürkle.
This is how the connection between the methodology and the field of application was first established. It was shortly thereafter, that Annika Krüger, a graduate of the Life Science study programme, started to conduct her doctoral thesis work. Her research project was funded through a scholarship from the Konstanz Research School Chemical Biology (KoRS-CB) and the Zukunftskolleg. It was during this time that Annika Krüger developed a customized procedure to investigate protein-nucleic acid interactions, using IR spectroscopy to observe structural changes in the protein.
Documenting structural changes
Implementing this procedure was not that easy at the beginning: the p53 protein is extremely complex. In order to investigate the flexible protein structures in the p53 molecule via IR spectroscopy, existing measurement procedures had to be adapted and further developed. The aim was to avoid the proteins’ unspecific interactions during the measurement process and to specifically bind its interaction partners to a surface. By carrying out these modifications, Annika Krüger was able to take advantage of IR spectroscopy’s tremendous sensitivity for the purpose of investigating the smallest changes in protein structures.
The doctoral candidate recognized the potential of this measurement technique compared to conventional structural analytical methods, since it is especially suitable for capturing the dynamic structural domains of the p53 protein. These intrinsically disordered –and still largely unexplored – regions within the protein play a central role in the regulation of p53, explains Annika Krüger: “We no longer have to exclude these dynamic regions, and instead, can observe both how they are regulated and the effects they have on the binding behaviour and the overall structure of the protein”.
The interplay between molecules
The research team’s spectroscopic approach also allowed it to learn more about the mechanisms involved when molecules interact. In order to better understand the different binding mechanisms utilized in the cellular stress response process, the Konstanz-based researchers investigated the interaction between the p53 protein and both DNA and biopolymer poly(ADP-ribose), in short PAR.
PAR plays an important role in the cellular stress response to DNA damage. When DNA is damaged, PAR forms in the cell nucleus and serves as a signal in the coordination of the cellular stress response. This process also regulates, for example, the protein activity of p53. Additionally, the researchers wanted to find out if the p53 protein changes its structure and possibly its function once it binds to PAR. “That PAR can strongly bind with p53 was already known when we started our research project. The question that previously did not have an answer, however, was ‘what happens to the protein on the structural level?’,” explains Aswin Mangerich. Using their enhanced approach, the researchers found out that p53 does not bind the same way to PAR as it does to DNA. As a result of observing small but clear structural differences, Annika Krüger concluded that p53 utilizes two different binding mechanisms, which may also play an important role in the regulation of p53. The research team’s efforts clearly demonstrate how their methodological approach can provide detailed insights into cellular processes.
The study’s authors agree: the close collaboration between researchers in the fields of biology and chemistry at the University of Konstanz proved to be a decisive factor in carrying out their successful research. Their interdisciplinary project and their joint effort in further developing their research techniques allowed them to take a close and unique look at the flexible domains of a protein and their functions. These new insights may help to create a more solid empirical foundation for future studies in the research fields of cancer and ageing.
Original publication: Krüger, A., Stier, A., Fischbach, A., Bürkle, A., Hauser, K., Mangerich, A. "Interactions of p53 with poly(ADP-ribose) and DNA induce distinct changes in protein structure as revealed by ATR-FTIR spectroscopy", Nucleic Acids Research, 2019, 47, 4843–4858